 .
.Nevertheless, we must make important, even critical decisions about testing and treatments, and despite our uncertainties about the bases for those decisions, the decisions themselves must be definitive. The patient must choose whether to undergo the recommended surgical treatment, despite remaining doubts about whether it is truly necessary; the physician must decide what test to perform next, despite conflicting desires for various different test results. People are, in fact, often uncomfortable making decisions for which they understand the rationale to be uncertain, and they seek means to relieve these uncertainties. In medicine, for example, one typical approach is temporizing: putting off the final decision and hoping that additional information will evolve to make the decision more straightforward. Another is buying additional information by performing less risky or costly tests that help to reduce the uncertainties in the more critical decision.
This paper surveys historical and contemporary approaches taken by medical informatics to dealing with uncertainty and the need to make decisions in the fact of uncertainty. We begin with a characterization of the issues around which research is organized. We then review the probabilistic approach to inference under uncertainty and the decision analytic approach to reaching decisions under uncertainty. We conclude by briefly examining some interesting ideas that approximate or abstract away from these classical approaches.
Other papers discuss the applied psychology of human judgments of uncertainty. For example, one examines the move from psychiatric interviews to diagnostic decisions, another considers feedback from therapy preferences to diagnostic plans, several concern accuracy and bias in human experts' intuitive judgments, and another examines violations of "rationality." One paper argues that women health care providers are less directive than men in the face of uncertainty.
Another set of papers examines methods of expressing uncertainty based on other than numerical measures. For example, several look at uncertainty based on linguistic data, one pursues logical methods in medical reasoning, and one tackles the meaning of the "non-" prefix in nosology.
A few papers concern methods for estimating uncertainty in specific situations. For example, one studies information elicitation for an image understanding system using Bayesian methods, another compares risk assessment by judgment of risks vs. physiological basis of risk estimates, and another looks at the effect of uncertain paternity on pedigree analyses in animal lines. One concerns interobserver variation in endoscopic diagnosis.
Not surprisingly, the Medline search turned up a number of articles of this kind, including ones examining the treatment of asthma, diagnosis of Huntington's disease, acute myocardial infarction in patients with acute thoracic pain, the effectiveness of alternate forms of ECMO (extracorporeal membranous oxygenation), modes of inheritance in breast cancer, and improvements in planning of radiation treatment.
Despite their aim to reduce uncertainty, most articles of this kind naturally fall outside the scope of a bibliographic search on the word "uncertainty," and may never actually mention the word. For authors of such papers, the desire to reduce uncertainty is tacit and so obvious that it is "beneath mention."
Two of the articles consider how well patients understand their risks and options. One concerns patients' attitudes and expectations in facing genetic counseling. The other examines patients' understanding of the specific risk factors and their consequences in two genetic disorders, retinitis pigmentosa and choroideremia.
Interestingly, four papers discuss the negative impact of the anxiety caused by uncertainty itself; it may affect the patient's mood and willingness to cooperate. One pursues this topic in the context of nursing for arthritis patients. Another studies the effect of using euphemisms for describing their disease to patients with cancer. Another discusses truth-telling among American doctors in the care of dying patients.
Efforts to quantify uncertainty in specific medical domains have typically been treated as a separate problem, to be solved by others, though many of the early projects in AIM [22,23,35] did in fact build substantial knowledge bases that included pseudo-probabilistic models of uncertainty (vide infra). The important issues of how patients and doctors deal with the psychological stresses of uncertainty have not received much attention, though preliminary studies such as [10] point to interesting ways of incorporating such concerns into a program's interface.
Medical informatics has changed tremendously over the past few decades, and changes in the approach to uncertainty are probably not the most important advances in the field. In [37] we discuss some of these other important changes. Perhaps the most important is the envisioned role of computer programs in health care. Twenty years ago most researchers expected programs to play the role of expert consultants, sought out by practicing physicians and nurses to evaluate and analyze difficult cases. Today, we instead expect background use of programs to detect potential errors and to recognize treatment opportunities, coupled intimately to evolving comprehensive medical record systems.
This point of view is somewhat controversial, because it does assign a primary role to probability theory. The issues are similar to those discussed in the field of artificial intelligence, where many researchers propose a similar role for predicate logic [7].
Given an a priori probability distribution over a set of possible states of the world (e.g., alternative diagnoses), and a conditional probability distribution telling how the outcome of a test depends on the states of the world, we can use the Reverend Bayes' theorem(1) to compute the a posteriori probability after we know the outcome of the test. Let D1, ... , Dn be the possible states of the world. We assume that they are exhaustive (i.e., no other states are possible) and mutually exclusive (i.e., only one state can hold). These assumptions assure that
The assumption of exhaustiveness requires that the model be complete for the population of problems to which it is to be applied. Violations of this requirement can lead to nonsense results, because---by definition---one of the possible hypotheses must be correct. Unless "other" is a valid hypothesis, a problem inappropriately brought to a program assuming exhaustiveness of its hypothesis set will be inappropriately diagnosed. But "other" is a difficult hypothesis to characterize---what is its prior likelihood, and what are its predicted manifestations? The second assumption, that only one state can hold, is only mildly troubling if only acute disorders are considered. This is because it is actually rather unlikely that two independent acute diseases will strike an individual at the same time. If chronic disorders are also under consideration, however, then the assumption of mutual exclusivity is often erroneous [21], and will tend to make it difficult to identify all of a patient's problems.
The conditional probability of a symptom S given D is simply the probability that S occurs when D occurs. We write  for the conditional probability of S given that Di is the true state of the world, and this is equal to
for the conditional probability of S given that Di is the true state of the world, and this is equal to  . Bayes' rule is then:
. Bayes' rule is then:
where by the closed-world assumption that S must be caused by one of the Di,
When using Bayesian inference where there are many possible independent observations (symptoms) possible, it is common (though not necessarily correct) to assume conditional independence. Mathematically, two symptoms, S1 and S2 are conditionally independent just in case
Intuitively, the symptoms are conditionally independent if they are linked only through the diseases that cause them, but are otherwise unrelated.
If we observe two different symptoms, it is possible to apply Bayes' rule once to calculate the posterior probability distribution for diseases by using this last formula. This is equivalent to treating the two symptoms together as a single compound symptom, and using that formula to compute the conditional probability of the compound symptom given each disease from the conditional probabilities of the individual symptoms alone. Of course if we have reason to believe that the symptoms are not conditionally independent, we can directly estimate the joint conditional probabilities, which are in that case no longer just the product of the individual ones. Note, however, that if many subsets of a large number of symptoms are conditionally dependent, we must estimate a vast number of joint conditional probabilities.
Instead of forming a single compound symptom from all the ones observed, we can adopt a sequential Bayesian inference method [8]. After observing any symptom, say Sl, we use Bayes' rule to compute the posterior probability  . Then, we may treat that posterior as a new prior probability, corresponding to the likelihood that the patient has Di if he is selected from a population of people all of whom have the symptom
. Then, we may treat that posterior as a new prior probability, corresponding to the likelihood that the patient has Di if he is selected from a population of people all of whom have the symptom  . We then repeat the same steps for all other observed symptoms, at each step using the posterior probabilities as the new priors for the next step. If conditional independence holds, we can show that this is equivalent to computing conditionals for the compound symptoms, as suggested above. Its advantage is that it allows us to consider doing the diagnostic reasoning step by step, and therefore to reason between steps about which would be the most useful symptom to try to observe next.
. We then repeat the same steps for all other observed symptoms, at each step using the posterior probabilities as the new priors for the next step. If conditional independence holds, we can show that this is equivalent to computing conditionals for the compound symptoms, as suggested above. Its advantage is that it allows us to consider doing the diagnostic reasoning step by step, and therefore to reason between steps about which would be the most useful symptom to try to observe next.
Especially in sequential application, it is very convenient to reformulate Bayes' Rule to use odds likelihood ratios rather than direct probabilities. If we compute not only  but also
but also  , it is clear that they share the same denominator. If instead of asking the posterior probability of D, we ask for the posterior odds of D, we have the odds-likelihood form of Bayes' Rule:
, it is clear that they share the same denominator. If instead of asking the posterior probability of D, we ask for the posterior odds of D, we have the odds-likelihood form of Bayes' Rule:
where the prior odds of D is given by
and the conditional odds (or likelihood ratio) of the symptom given the disease is given by
Incidentally, there is something potentially troublesome in a term such as  when D is not a binary variable. If D is k-ary, then it must take on one of the values
when D is not a binary variable. If D is k-ary, then it must take on one of the values  . If, say,
. If, say,  is the distinguished value whose absence we denote by
is the distinguished value whose absence we denote by  , then
, then  must correspond to some superposition of the states in which D has the values
must correspond to some superposition of the states in which D has the values  . But then
. But then  . This expression is not, in general, a constant, but will vary with the relative likelihoods of the other possible values of D. Assuming it to be a constant is, in fact, an additional assumption that does not follow from the more typical assumption of conditional independence, but a number of papers in the literature do not appreciate this.
. This expression is not, in general, a constant, but will vary with the relative likelihoods of the other possible values of D. Assuming it to be a constant is, in fact, an additional assumption that does not follow from the more typical assumption of conditional independence, but a number of papers in the literature do not appreciate this.
This formulation leads us to talk about "three-to-one odds" instead of "75% probabilities", but has the advantage that successive applications of Bayes' rule consist merely of successive multiplications by the likelihood ratios corresponding to successive observations, if those observations are conditionally independent. A further advantage of the odds-likelihood notation is that people untrained in probabilistic reasoning appear to be somewhat better at being able to estimate odds corresponding to very small and very large probabilities than at estimating the probabilities themselves---though this is a controversial claim. For example, the difference between a 98% or a 99% probability of some event seems small to many people, but actually corresponds to a difference in odds between 50:1 and 100:1.
When conditional dependence is present, we need to use joint likelihood ratios, in exact analogy to joint conditional probabilities. For example, if  and
and  are not conditionally independent given D, then we must directly estimate
are not conditionally independent given D, then we must directly estimate  , which will not, in general, equal
, which will not, in general, equal  .
.
Pearl [25] points out a very important conceptual use of the likelihood ratio in interpreting uncertain evidence. When evidence is unreliable, the best way to express its significance may be to judge how much more likely it is in the presence of an hypothesis than in its absence. Pearl imagines a drunken neighbor's report of a burglar alarm sounding; even though it is difficult to model just how the neighbor's drunkenness and the alarm sound might interact to cause him to make this report, we may summarize its effect by saying that it is twice as likely to occur if the alarm is in fact sounding than if not. This corresponds to conditional odds of 2:1, which can be used directly in (5) to update the likelihood of the alarm having sounded (and ultimately of a burglary having taken place).
If the odds form of Bayes rule is transformed by taking the logarithm of both sides of the equation, we arrive at a formula that computes the log odds of a hypothesis as the sum of the log likelihood ratios of the observed evidence. In this formulation, each piece of evidence contributes a weight of evidence to the hypothesis. Findings that have a likelihood ratio greater than one (i.e., that are more likely to be found if the hypothesis is correct than if not) will contribute a positive weight, whereas those with a likelihood ratio less than one contribute negative weight. Equivocal evidence, which is neither more nor less likely depending on the presence of the hypothesis, contributes nothing.
Many early AIM programs in fact used additive scoring techniques to estimate the likelihood of various hypotheses. The frequency measure used in Internist [22], for example, is best interpreted as a scaled log likelihood ratio.
The approach summarized above is the core of most probabilistic reasoning systems. The restrictive assumptions make efficient computation possible and make it relatively easy to acquire the needed probabilities, but also make the models unrealistic for many real-world problems.

 . Given the prior probabilities of the Di and these relevant conditional probability tables, we may continue to apply a method very much like Bayes' rule to compute the probabilities of each of the Di after having observed any of the Sj. For large numbers of diseases and symptoms, however, both the sizes of the conditional probability tables and the computation needed grow exponentially and become impractical.
. Given the prior probabilities of the Di and these relevant conditional probability tables, we may continue to apply a method very much like Bayes' rule to compute the probabilities of each of the Di after having observed any of the Sj. For large numbers of diseases and symptoms, however, both the sizes of the conditional probability tables and the computation needed grow exponentially and become impractical.

This assumption is legitimate only if there is no special reason to believe that particular combinations of causes of a symptom make it more or less likely. If appropriate, it reduces the amount of data needed to characterize a conditional probability table for a symptom with k possible causes from 2k to k, a very significant saving.
There is actually a subtle problem in the formulation of noisy-or above, having to do with how we estimate conditional probabilities like  . The probability that some disease causes a symptom is not generally equal to the probability that the symptom is present if the disease is. The latter will generally be a higher number, because it is possible that some other cause of the symptom was also present and actually caused the symptom. When we estimate causal probabilities, we really mean "the probability that S1 occurs given that D1 is present, but no other diseases are." This is
. The probability that some disease causes a symptom is not generally equal to the probability that the symptom is present if the disease is. The latter will generally be a higher number, because it is possible that some other cause of the symptom was also present and actually caused the symptom. When we estimate causal probabilities, we really mean "the probability that S1 occurs given that D1 is present, but no other diseases are." This is
The noisy-or approximation then applies (8) using these causal probability estimates for the conditional probabilities.
Another complexity that arises in models such as these is that, because of known but unavoidable incompleteness of the model, a symptom may in fact occur in the absence of any of its possible causes. A simple but apparently adequate approach to this problem has been to estimate an additional (constant) leak term for the noisy-or, which represents the probability of the symptom arising from the everpresent but unmodeled background.
Noisy-and is a similar modeling assumption, corresponding to situations in which all of a symptom's causes are necessary for it to appear. For example, a person must be both susceptible to and exposed to a pathogen for its effects to be present; even if both are true, the effect may still fail to arise. There have been proposals for additional simplifying models for how multiple causes interact to provide the probability of a joint effect, though they are used far less often than noisy-or. The most interesting correspond roughly to the causal models of [29].
There is an important special case of these topologies, wherein there is at most one path between any two nodes in the network---such networks are called polytrees. This condition assures that any prior knowledge or observation can influence any other node via but one path. Then, a slight generalization of the sequential updating methods described above will successfully calculate the revised likelihood of any node in a time proportional to the diameter of the graph [25], which is very good. Polytrees provide an important and useful class of models, because they permit the aggregation of multiple pieces of evidence, "causal" chains of reasoning, and "explaining away" evidence, yet can be computed efficiently. However, these polytree methods fail as soon as we draw additional links in the graph, making multiple pathways for evidence propagation possible.
An appropriate formulation of the probabilistic inference problem in this general case was only worked out in the early 1980's, mostly by Pearl and his students [25], though specialized versions for pedigree analysis were understood by geneticists by the mid-70's [6,19].
Consider a small example of a network of this type, shown in Fig. 3, where for simplicity we will assume that each node is binary: it is either present or absent.

We can easily compute the joint probability of any particular set of values for the five nodes. For example, the probability that all five are present is given by
Similarly, the probability that A, C and D are present but B and E are absent is
In general, if the nodes of a network are Xi and the set of parent nodes of X is  , then the probability of any particular set of values for all the nodes in the network is given by
, then the probability of any particular set of values for all the nodes in the network is given by
For nodes with no parents,  should be read as just
should be read as just  , the prior probability.
, the prior probability.
Usually, we are not interested in the probability of some state of the world in which all nodes have specific values, but in a partial state of the world in which only some nodes have specific values. We might want to know, for example, what is the probability that the patient has some specific disease even when we do not know the presence or absence of all symptoms, syndromes and other diseases. In this case, we must simply sum over all possible states of those nodes whose value we do not know. Thus, if we do not know the values of the first k of the n nodes, then
The sum is taken over all possible combinations of values of the nodes  , and each term of the sum may be computed by (12).
, and each term of the sum may be computed by (12).
In the simple example above, the probability that A is present and E is absent (over all possible states for B, C, and D) is thus given by
and the summation is over the eight terms corresponding to all possible combinations of values of B, C, and D. By (10), each term requires looking up five conditional probabilities and multiplying them together. Thus, we will perform 8x4=32 multiplications if the computation is done naively.
This version of the calculation requires only 9 multiplications, because it recognizes facts such as that the dependence of  on A through C does not depend on B or D. Such savings become spectacular for very large networks, if the factorization is done well.
on A through C does not depend on B or D. Such savings become spectacular for very large networks, if the factorization is done well.
Most approaches to solving this problem have viewed it not as a symbolic algebra problem (as presented above) but as a graph-separation problem [18]. Most algorithms attempt to find small subsets of the original graph whose nodes, if known, render other large subsets independent. For example, in MCBN1, above, knowing the value of C assures that the probability of E does not depend on the values of A, B, or D, and vice versa. This observation corresponds exactly to the factorization of the algebraic formula in (15), and the correspondence is universal: for every good graph separation there is a good factorization, and vice versa.
Unfortunately, most interesting problems of interest about the general case of Bayes networks are NP-hard, including exact solution of a network and optimal structuring of the computations---optimally factoring a large expression or finding optimal graph separators [4]. Thus, the best algorithms are heuristics that seem mostly to do a very good job, but that do not guarantee the best possible results.
Gaussian networks are an interesting special case of efficiently-solvable Bayes networks [32]. In these, instead of nodes taking on discrete values and probabilities being described by conditional probability tables, we assume that all nodes take on continuous values whose probabilities are normally distributed around a mean, with some standard deviation. Conditional probability distributions are, then, multidimensional normals, and it turns out that polytree-like propagation algorithms that adjust the means and standard deviations of their neighbors rapidly solve the inference problem.
 , then say A is present, otherwise absent.
, then say A is present, otherwise absent.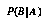 or
or  , depending on whether A was assigned + or --.
, depending on whether A was assigned + or --.The evaluation of large Bayes networks is an area of much current research (e.g., [33]), and many groups are exploring useful applications. Only some are diagnostic. For example, several groups in natural language processing use Bayes nets to disambiguate word senses, parse spoken speech, and predict which journal articles are likely to be of interest to a person based on the previous articles in which they have shown an interest.
One of the simplest ideas in this vein is to update the likelihoods in an existing structural probabilistic model by adjusting the conditional (and maybe the prior) probabilities based on actual data. Spiegelhalter [36] suggests indicating the model builder's confidence in a probability by expressing it as a ratio of the number of times the indicated event would occur in a number of trials. Thus, a conditional likelihood of 1/2 has the same numerical value as one of 500/1000, but shows far less confidence in the number. Then, actual data from subsequent experience with the system can be used very directly to update the probabilities. One negative sample would change the initial 1/2 estimate to 1/3, but it would change the more confident 500/1000 estimate only to 500/1001, an almost negligible change. As the amount of actual data increases, the model's initial probability estimates are adjusted toward and eventually supplanted by estimates from the data. Although this is very desirable, problems of selection and ascertainment can nevertheless corrupt the model.
More dramatic opportunities are explored by systems that can learn not only the parameters of a Bayesian network but its structure as well. Pearl and Verma [26] exploit statistical evidence of independence among random variables and asymmetries in the independence relations among triples of variables that are causally linked to identify plausible causal structures from empirical evidence. Cooper and Herskovits [4] define a heuristic search algorithm to find the most probable Bayes network structure given a set of data, and show that they can quite reliably recover Bayes networks from data sets generated by statistical sampling from those networks.
Another interesting approach to the problem of learning Bayes networks strives to improve the efficiency of knowledge acquisition not by learning from data but by making it easier for experts to express their judgments. Heckerman introduced probabilistic similarity networks that allow the expert to concentrate on the individual differentiation problems that are crucial to a domain. His system then builds a complete Bayes network by aggregating the locally-expressed judgments [13].
A decision tree is typically (though not necessarily) ordered so that the root of the tree is the first decision that must be made, and further down each branch may come additional decisions that arise depending on previous decisions and chance results. Decision trees can, therefore, represent contingent plans, because later decisions may become relevant only if chance outcomes lead to them.
The decision tree in Fig. 1 represents the sequence of decisions faced by a patient who has a very badly infected foot that has turned gangrenous. The immediate choices are to amputate the foot or to treat medically. Depending on the outcome of the medical treatment, the patient may face the choice of subsequently having to amputate the whole leg or continuing with medical treatment. For the (fictitious) numbers used in the figure, the best decision is to treat medically, but then to amputate the leg if the gangrene gets worse after medical treatment. Normally, one would not simply make such a conclusion and act on it, because both the probability estimates and the utilities are tenuous. Instead, one would typically undertake a sensitivity analysis---to see whether the decision is robust against small changes in various parameters. For example, how much would we have to raise the value of life without a foot before an immediate foot amputation would appear preferable to medical treatment and the risk of losing a whole leg. How much more likely would death after initial medical treatment need to be before we (or the patient) would prefer immediate amputation? Is the decision sensitive to the estimate of the utility of recovery after two courses of medical treatment? In the real world, such sensitivity analyses often suggest that critical additional data be gathered to help resolve the uncertainty.

Influence diagrams are considerably more compact representations of large decision problems than decision trees, because they do not explicitly expand the consequences of individual choices or chance occurrences. Instead, each arc carries a matrix of probabilistic relationships between all possible states of the source and target of the arc. Influence diagrams are equivalent to fully symmetric decision trees, and this can be both an advantage and disadvantage. It assures that no unintended asymmetry arises in the analysis of a complex decision problem. However, it makes it impossible (within the representation) to simplify the model by omitting branches that have very low probability or that don't significantly influence the utility model. Some practitioners also find the more concrete representation of the decision tree easier to understand. Because influence diagrams and decision trees are interconvertible, it is also possible to build hybrid modeling tools that permit viewing a decision problem from either vantage point.
The classic problem in reasoning about individual utilities is that they are so difficult to quantify. According to decision theory, standard lottery techniques must be able to determine a numerical utility scale for all possible circumstances for a rational decision-maker, assuming only very reasonable constraints. Thus, if we assume that the ends of the utility scale are defined by the state of good health (worth 1000, in our amputation scenario) and death (0), then any other intermediate state can be assigned a value in the following manner: Offer the decision-maker a choice between that state for certain or a gamble in which the probability of getting good health is p and the probability of death is (1-p). If we vary p, there should be a value p* for which the decision-maker is indifferent to whether he chooses the certain outcome or the gamble. Then, 1000*p is the value of the state in question. Despite the attractiveness of this procedure, people find it difficult to make such judgments reliably, they produce judgments that differ significantly depending on just how the question is phrased, and they may be unable to estimate the value of a hypothetical state until they are actually faced with the decision. We address the consequences of some of these problems near the end of the current paper, but unfortunately there are no satisfactory general solutions to this problem.
In applying decision analysis to clinical medicine, doctors have found one instance of the utility problem to be quite common: many decisions may affect the complete future course of a patient's life, so there is no short time horizon over which one can assess the benefits and defects of a course of action. For example, a decision to perform coronary bypass surgery now will affect the patient's quality of life for many years to come, and will also alter the expected likelihood of various contingencies of interest such as restenosis. To model the value of a current intervention thus requires modeling all its possible long-term consequences. One successful approach to this problem is to assume that the future course of a patient's state is determined by a Markov process, and that the intervention sets the parameters of that process [1]. If each state has a given value, then the long-term value of a decision is calculated by integrating the value of each state times the probability of being in it over a long enough time to encompass the maximum expected life of the patient. This approximation assumes that the critical decision itself is made now, and the rest of the analysis merely explores its consequences over time.
Berzuini et al. [2] construct general Bayes networks whose structure reflects repeating episodes, each depending on a set of global patient and process-specific parameters. The beauty of this approach is that the usual inference mechanisms of Bayes networks serve both to estimate the parameters from observed episodes and to predict future behavior of the system.
More general temporal models currently being explored consider cases in which there is not a single decision to be made but a recurring sequence of decisions, in which the optimal strategy may well be to make different decisions in the future than at present. Leong [20] casts the problem in the framework of semi-Markov decision models and is building tools to support construction and evaluation of such models.
Early AIM and AI programs introduced a large variety of scoring schemes that were thought, at the time, to be simpler or more attractive than probability theory. In retrospect, however, many of these have been shown to be equivalent to standard probability theory, with perhaps a few additional assumptions or approximations (e.g., [12] concerning Mycin, and the discussion of log likelihood ratios, above, for Internist). Some of the schemes were originally introduced simply because the methods of Bayes networks were unknown, yet the need for chains of probabilistic inference was critical (e.g., the inference scheme of Prospector [5]). The evolution of the powerful Bayes network formalism has obviated the need for many such scoring schemes, and has provided at least a background theory against which to measure any other proposals.
Abstraction that completely disregards numerical knowledge and exploits only the structural relationships can be quite powerful and easy to use. Reggia et al. developed a set covering diagnostic model [28] that uses the bipartite graph representation but without regard for actual probabilities. A diagnostic solution in this sense is a minimal set of diseases that can cause all of the observed symptoms. By Occam's razor, the simplest explanation of a problem is the best, and the set cover algorithm equates simplicity with the smallest number of hypothesized disorders. This corresponds roughly to assuming that every disease is equally likely, that all symptoms combine causal influences on them by noisy-or, and that the disease-to-symptom link strengths are all equal. In a domain rich in interconnections and alternative explanations of many phenomena, and when the amount of noise in both the models and observations is low, this can be an effective and efficient diagnostic scheme. Improved heuristic search methods and representations for partial solutions can gain further dramatic efficiency improvements [38,39].
People obviously do not use full-blown decision analysis to make decisions about their daily lives, yet they often act in quite rational ways. This suggests that there are many as yet unexplored abstractions and approximations of our formal models for reasoning under uncertainty that can be fruitful. They should lead to models that are easier to understand because they correspond to important aspects of the ways in which people think about problems. They may also be easier to compute than the full-blown, often exponential models that currently arise in exact calculations of results.
Nevertheless, these anticipated trends will create great opportunities for interesting research in medical reasoning under uncertainty. For the first time, we will move from problems of data scarcity to those of data overload, as the volume of recorded data explodes. Most of those data will be captured to satisfy regulatory and reimbursement needs, not primarily the needs of clinical care or of medical research. Therefore, tremendous opportunities will arise to develop insightful new approaches for taking advantage of this developing glut of data, helping to relate it to clinical and research needs and to the goals and preferences of tomorrow's patients.